ClariTest® Core Non-Invasive Prenatal Screening

ClariTest® Core is a non-invasive prenatal screen (NIPS) that identifies the risk for fetal chromosomal abnormalities. ClariTest Core can be performed as early as 10 weeks gestation from a simple blood draw. Results are available within five to seven days. ClariTest Core can be used to screen singleton and egg donor/IVF pregnancies for the common trisomies, sex chromosome aneuploidies and 22q11.2 microdeletions. Twin gestations can be screened for the common trisomies and for presence of the Y chromosome.
Conditions Included in ClariTest Core Screening
Click on a condition below to learn more
- For more information related to conditions or services, click here.
References for above conditions
- NIH U.S. National Library of Medicine: Down Syndrome
- National Down Syndrome Congress
- NIH U.S. National Library of Medicine: Trisomy 18
- NIH U.S. National Library of Medicine: Trisomy 13
- NIH U.S. National Library of Medicine: Turner Syndrome
- U.S. Departments of Health & Human Services: Turner Syndrome
- U.S. National Library of Medicine: Klinefelter Syndrome
- U.S. National Library of Medicine: Triple X Syndrome
- U.S. National Library of Medicine: 47,XYY Syndrome
- U.S. National Library of Medicine: 48,XXYY Syndrome
- McDonald-McGinn, D. M. & Sullivan, K. E. (2011). Chromosome 22q11.2 deletion syndrome (DiGeorge syndrome/velocardiofacial syndrome). Medicine, 90(1), 1–18.
Why screen for 22q11.2 microdeletion?
ClariTest Core offers the option to add screening for 22q11.2 microdeletion syndrome, also known as DiGeorge syndrome. DiGeorge syndrome is an autosomal dominant condition, and is the second most common genetic cause of heart defects and developmental delay, after Down syndrome.1 Unlike trisomies, maternal age does not increase the chance for 22q11.2 microdeletion, and more than 90% of affected individuals have no family history of the condition. 2
22q11.2 microdeletion affects as many as 1 in 1,000 pregnancies.3
The performance of the ClariTest Core prenatal screen for 22q11.2 microdeletion has been evaluated in the largest 22q11.2 validation study with over 1,900 hundred cases, 129 with confirmed deletions.4
Clinical Indications for ClariTest Core
- Routine pregnancy screening
- Advanced maternal age
- Positive maternal serum screen
- Abnormal ultrasound findings
- History suggestive of increased risk for common chromosomal abnormalities
ACMG recommends informing all pregnant women, including women at low or average risk, that NIPS is the most sensitive screening option for trisomy 13, 18, and 21.5 ACOG and SMFM confirm that cfDNA screening (NIPS) is an appropriate choice of testing for all pregnant women, regardless of age or risk and that NIPS is the most sensitive and specific screening test for common aneuploidies.16
Proven Technology
ClariTest Core utilizes the DANSR™ platform and FORTE™ algorithm, a widely studied cell-free DNA methodology, with over 74 peer-reviewed publications. The studies demonstrated exceptional performance in singleton and twin pregnancies and in women of any age or risk category6-13 and established the superior accuracy and reproducibility of the platform for fetal fraction assessment.14
| Disorder | N | Sensitivity | Specificity |
| Trisomy 212 | 23,155 | 99.3% (418/421) | 99.96% (22,724/22,734) |
| Trisomy 182 | 22,399 | 97.4% (147/151) | 99.98% (22,243/22,248) |
| Trisomy 132 | 14,243 | 93.8% (30/32) | 99.98% (14,208/14,211) |
| Monosomy X3 | 1,381 | 94.3% (82/87) | 99.8% (1,291/1,294) |
| 22q11.2 Microdeletion4 | 1,953 | 75.2% (97/129) | 99.6% (1,816/1,824) |
| XXX/XXY/XYY/XXYY3 | Other sex aneuploidies will be reported if detected. Limited data of these more rare aneuploidies preclude performance calculations. |
Fetal Sex: Accuracy for fetal sex (male or female) is >99%3
ClariTest Core utilizes microarray quantitation and a proprietary, targeted technology that includes both the DANSR assay and the FORTE algorithm to provide exceptionally accurate results.
DANSR: Digital Analysis of Selected Regions
- Specifically targets the chromosomes of interest for quantification (counting), providing targeted analysis
- Precisely measures and distinguishes fetal fraction using SNP technology
- Eliminates extraneous data from other chromosomes and unmapped cfDNA allowing for clear results
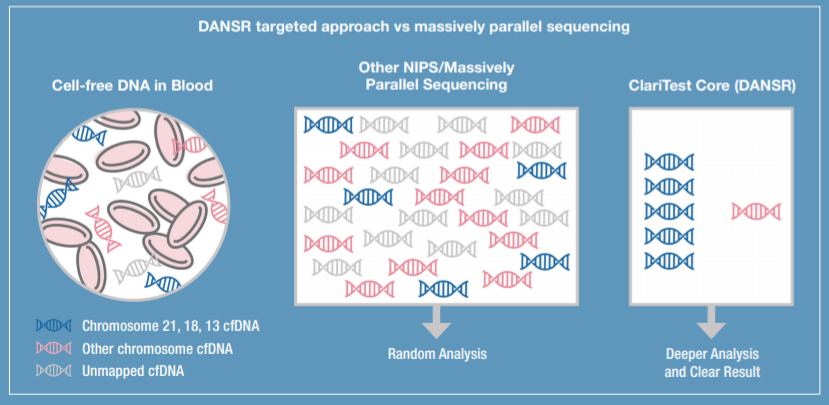
FORTE: Fetal Fraction Optimized Risk of Trisomy Evaluation
- Incorporates chromosome quantification and fetal fraction adding confidence to both high-and low-risk results across all fetal fractions
- Outperforms the Z-score approach (used by many NIPS platforms) regardless of the patient’s age or risk11
- Distinguishes high- and low-risk results with greater discrimination11
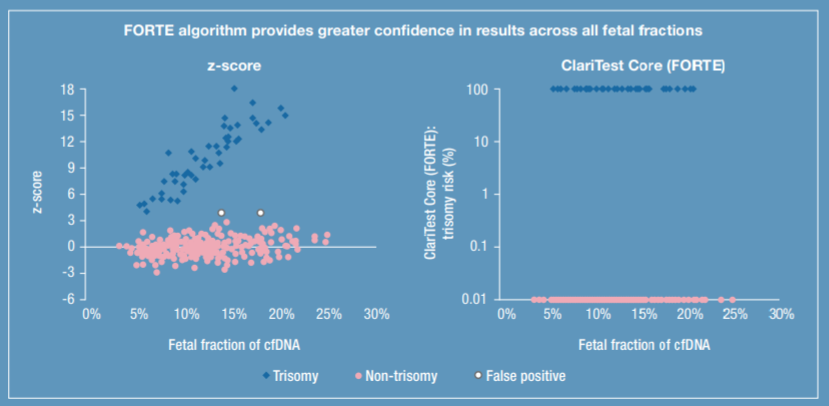
*z-score: a numerical measurement used in statistics of a value’s relationship to the mean (average) of a group of values, measured in terms of standard deviations from the mean.
Precise Results and Measurement of Fetal Fraction Using Microarray
- The DANSR assay products are quantified on a microarray platform. Incorporation of microarray technology results in increased precision in both result determination and fetal fraction measurement.15
- Utilization of microarray provides a three-fold increase in the number of SNPs assayed for fetal fraction compared to NGS, making the fetal fraction measurement more accurate and highly reproducable.15
- ClariTest Core NIPS requires a minimum 4% fetal fraction as a quality control standard, which ensures there is enough fetal DNA present to make a highly accurate call.
Clear, Accurate Reporting
- Easy-to-read reports with clear graphics to make triaging patient results faster
- High-risk results for Trisomy 21, 18, 13, Monosomy X or 22q11.2 Deletion syndrome include a positive predictive value (PPV)
- Low-risk results include a negative predictive value (NPV)
- Fetal fraction is included in the report
Results will be reported as high- or low-risk:
![]() High Risk: Positive Predictive Value (PPV) will be reported if result is high risk for trisomy 21, 18, 13, monosomy X, or a 22q11.2 microdeletion in singletons
High Risk: Positive Predictive Value (PPV) will be reported if result is high risk for trisomy 21, 18, 13, monosomy X, or a 22q11.2 microdeletion in singletons
![]() Low Risk: Negative Predictive Value (NPV) >99% for all conditions and is reported for low-risk singleton pregnancies
Low Risk: Negative Predictive Value (NPV) >99% for all conditions and is reported for low-risk singleton pregnancies
ClariTest Core is a screening test. Patients receiving a high risk result are advised to seek genetic counseling, as well asinc. to discuss diagnostic testing options. Decisions regarding the pregnancy should NOT be made based on ClariTest Core results only.
Insurance information
BioReference Health®, LLC and its division GenPath®, is an in-network provider for all major national and many regional health plans, increasing your access to ClariTest Core non-invasive prenatal screening.
References
- McDonald-McGinn DM et al. Nat Rev Dis Primers. 2015;1:15071. doi: 10.1038/nrdp.2015.71.
- McDonald-McGinn DM et al. Genet Med. 2001; 3(1):23-9.
- Grati FR et al. Prenat Diagn. 2015;35(8):801-809.
- Schmid M et al. Fetal Diagn Ther. 2018;44(4):299-304.
- Gregg AR et al. Genet Med. 2016;18(10):1056-65.
- Stokowski et al. Prenat Diagn. 2015;35(12):1243-6.
- Jones K et al. Ultrasound Obstet Gynecol. 2018 51:274-277.
- Norton ME et al. Am J Obstet Gynecol 2012;207:137.e1-8.
- Nicolaides KH et al. Am J Obstet Gynecol.2012 Nov;207(5):374.e1-6.
- Gil MM et al. Ultrasound Obstet Gynecol. 2019;53(6):734–742.
- Gil MM et al. Fetal Diagn Ther. 2014;35:204-11.
- Jones K et al. Ultrasound Obstet Gynecol. 2018;51(2): 275–6.
- Schmid M et al. Ultrasound Obstet Gynecol. 2018;51(6):813-17.
- Sparks AB et al. Am J Obstet Gynecol. 2012;206:319.e1-9.
- Juneau K et al. Fetal Diagn Ther. 2014;36(4):282-6.
- Rose NC et al. Am J Obstet Gynecol. 2020;136(4):e48-69.

