Solid Tumor NTRK Gene Fusion Assay
NTRK Gene Fusions are now established as an important biomarker for routine clinical oncology with pan-cancer, tumor-type agnostic implications for therapy and patient management. 1,2
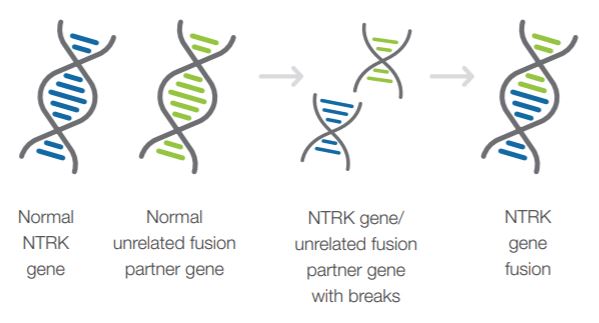
- NTRK Gene Fusion testing is a strongly recommended component of comprehensive genomic profiling of newly diagnosed solid tumors.1,3
- NCCN and other expert consensus recommendations indicate that NTRK gene fusion testing should be performed as part of a broad, panel-based approach.1
NTRK Fusions are identifiable among a wide range of common cancer types including NSCLC, GIST, CNS tumors and Melanoma, and are highly recurrent among particular rare cancer types. Recent, landmark studies have reported significant clinical responses to targeted NTRK inhibitors in up to 80% of NTRK fusion positive patients.4
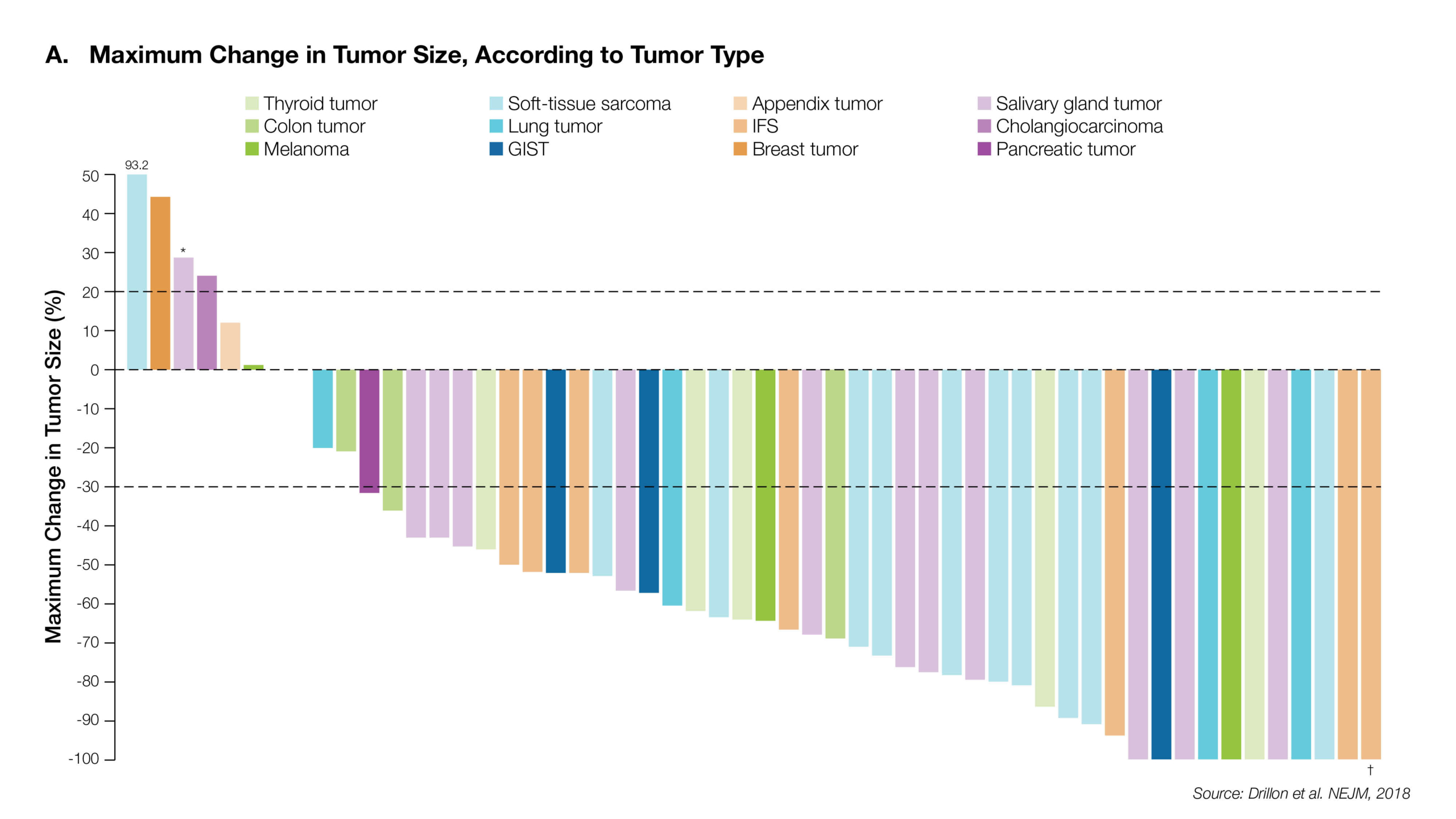
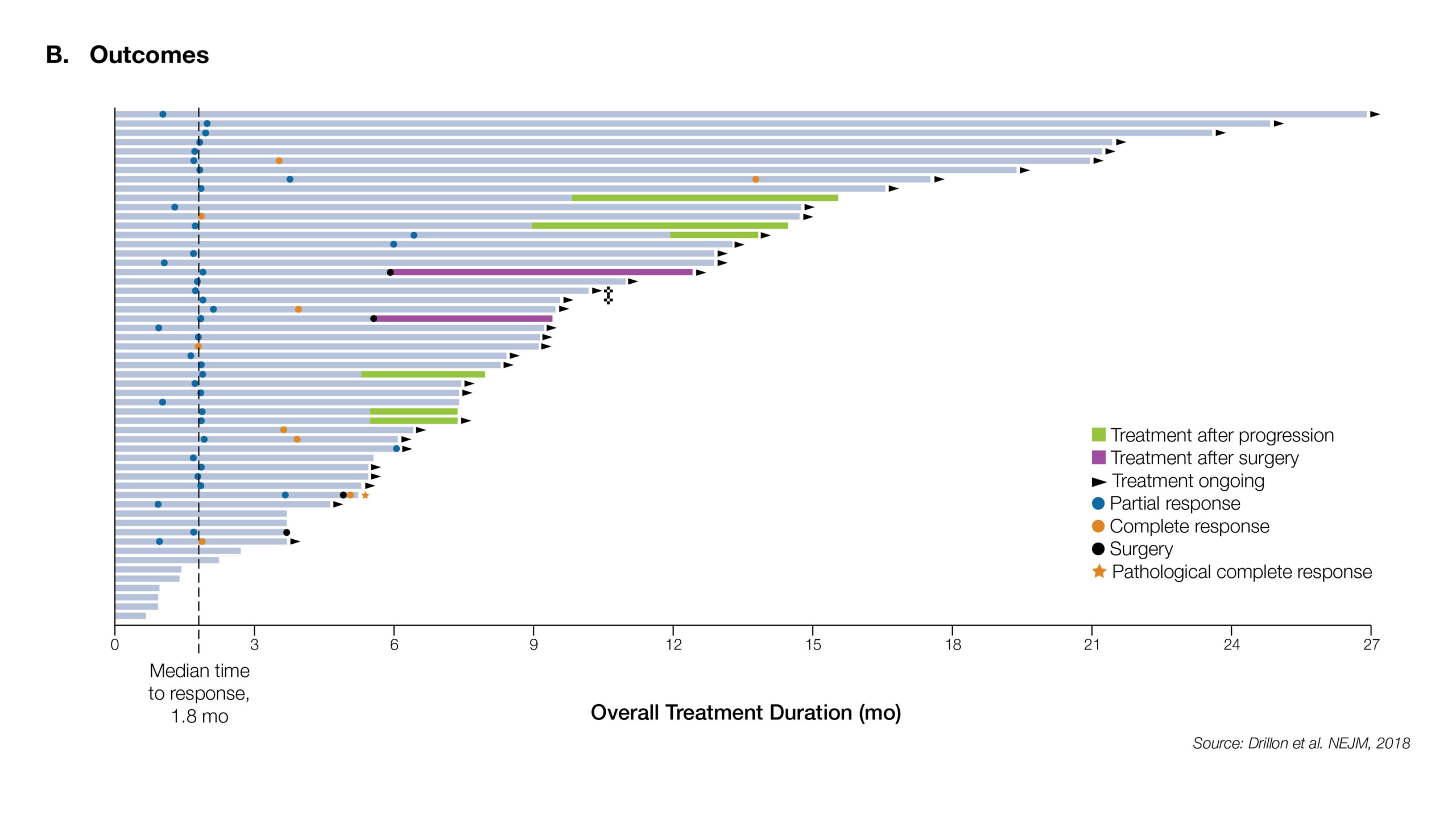
Estimated Frequency of NTRK Gene Fusion in Specific Tumor Types
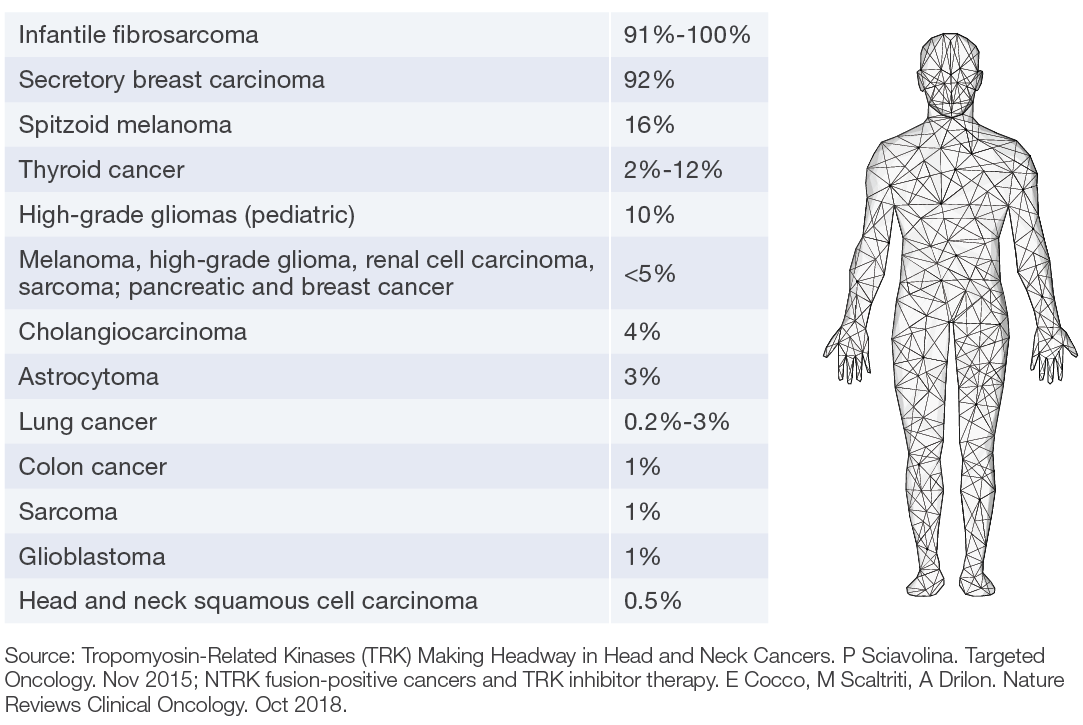
Advantages of GenPath NTRK Gene Fusion Testing
- RNA-based next-generation sequencing (NGS) is the technology of choice to definitively identify gene fusion events involving NTRK1, 2 or 3. RNA NGS fusion panel testing confers significant practical and technical advantages in comparison to traditional methodologies, and may limit risks false negatives or nonspecific results.1,3
- TAT is minimized. Turnaround time to results averages 7-10 days upon specimen receipt.
- Multi-gene RNA NGS fusion panels minimize tissue depletion in scant samples. RNA NGS provides a single test solution to simultaneously and comprehensively target broad ranges of gene fusions relevant to clinical oncology. Compared to traditional methodologies that may require application of multiple probes (e.g. FISH), risk for tissue depletion and QNS rates among often scant FFPE samples may be minimized (e.g. Non-Small Cell Lung Cancer needle core biopsies).3
- RNA NGS translocation detection is binding partner agnostic. Traditional methodologies (e.g. FISH, PCR) detect chimeric fusions involving only known, selectively targeted genes using pre-designed probes or primers. By contrast, RNA NGS readily delineates novel, previously unknown fusion partners or known binding partners that may be highly variable. Detection sensitivity is therefore significantly improved for gene fusion events that may be disease-defining or therapeutically predictive.
See sample report here.
| Test Code | Test Name | Methodology | Specimen Type | TAT |
| J355-9 | Solid Tumor NTRK Gene Fusion Assay (18 genes)
ALK, AXL, BRAF, CCND1, EGFR, FGFR1, FGFR2, FGFR3, MET, NRG1, NTRK1, NTRK2, NTRK3, PPARG, RAF1, RET, ROS1, THADA |
Next-generation Sequencing (NGS) | FFPE: Specimen Block Preferred Slides: 15 unstained slides cut at 5 microns containing a minimum 10% tumor cellularity | 7-10 days |
References
- Hsiao S.J et al Detection of Tumor NTRK Gene Fusions to identify patients who may benefit from Tyrosine Kinase (TRK) Inhibitor Therapy.
- Amatu A, SartoreBianchi A, Siena S. NTRK gene fusions as novel targets of cancer therapy across multiple tumour types. ESMO Open 2016;1:e000023. doi:10.1136/esmoopen-2015- 000023
- NCCN Guidelines, Non-Small Cell Lung Cancer, Version 3.2020.
- Drilon et al. Efficacy of Larotrectinib in TRK Fusion-Positive Cancers in Adults and Children. N Engl J Med. 2018 Feb 22;378(8):731-739

